QIIME 2 Enables Comprehensive End‐to‐End Analysis of Diverse Microbiome Data and Comparative Studies with Publicly Available Data - Estaki - 2020 - Current Protocols in Bioinformatics - Wiley Online Library
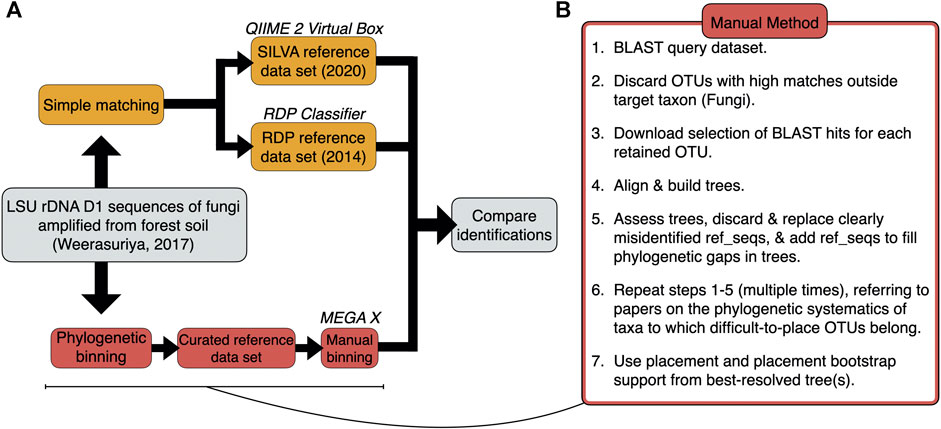
Frontiers | Simple Matching Using QIIME 2 and RDP Reveals Misidentified Sequences and an Underrepresentation of Fungi in Reference Datasets